Mnova
What's New in Mnova 17
Mnova 17 is now available with a refreshed user experience, smoother analysis, expanded scripting, plus upgrades across NMR, MSChrom, Dynamics, DB/Server and Gears, all shaped by real user feedback.
About Mnova
Mnova is a professional analytical chemistry software suite for processing, analyzing, visualizing, and reporting data from NMR, LC-MS/GC-MS, and other spectroscopic techniques within a single, vendor-neutral platform. Its modular design lets labs tailor workflows from routine QC to advanced research, while maintaining consistency across datasets.
Trusted by over 8,000 organizations worldwide, including 41 of the top 50 pharmaceutical companies, Mnova supports groundbreaking discoveries and unifies disparate data under one platform. In academia, it's the leading platform for NMR data processing, having analyzed millions of spectra to accelerate research and enhance outcomes globally. By reducing software fragmentation and minimizing tool switching, Mnova enables labs to work faster, smarter, and more efficiently, delivering reliable results across the entire analytical workflow.
Products
Features
One Interface for All Your Data
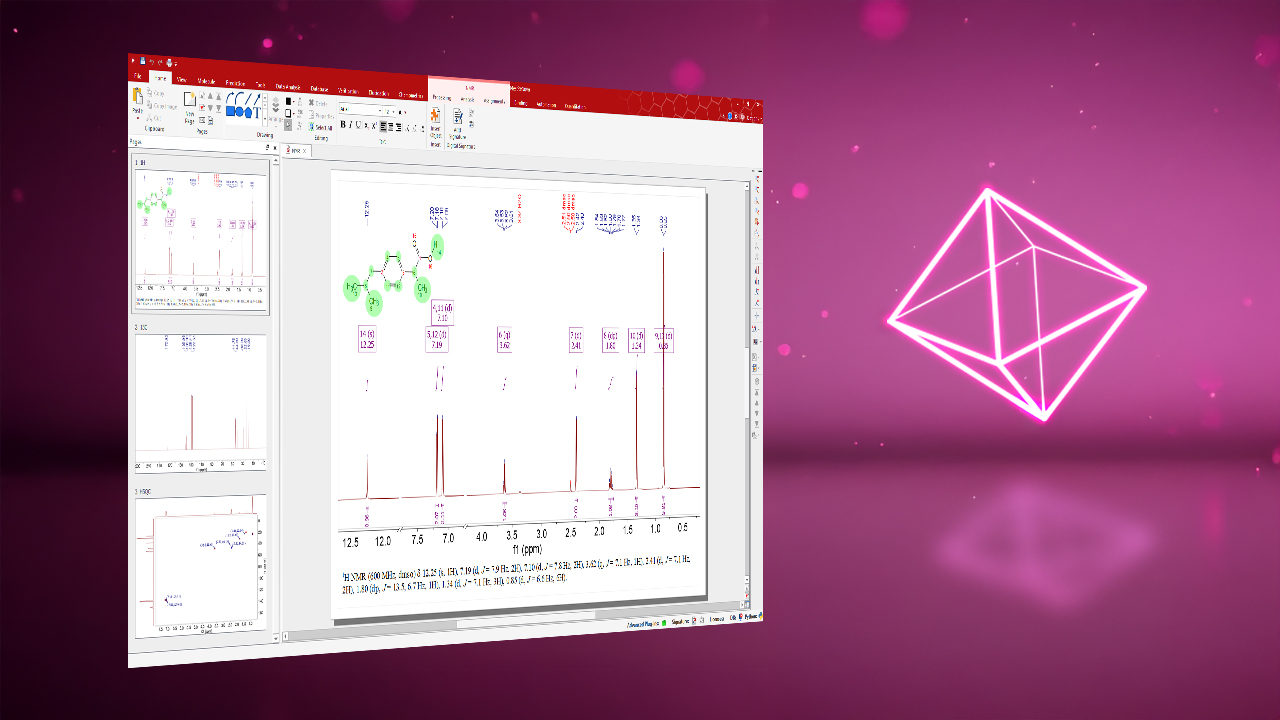
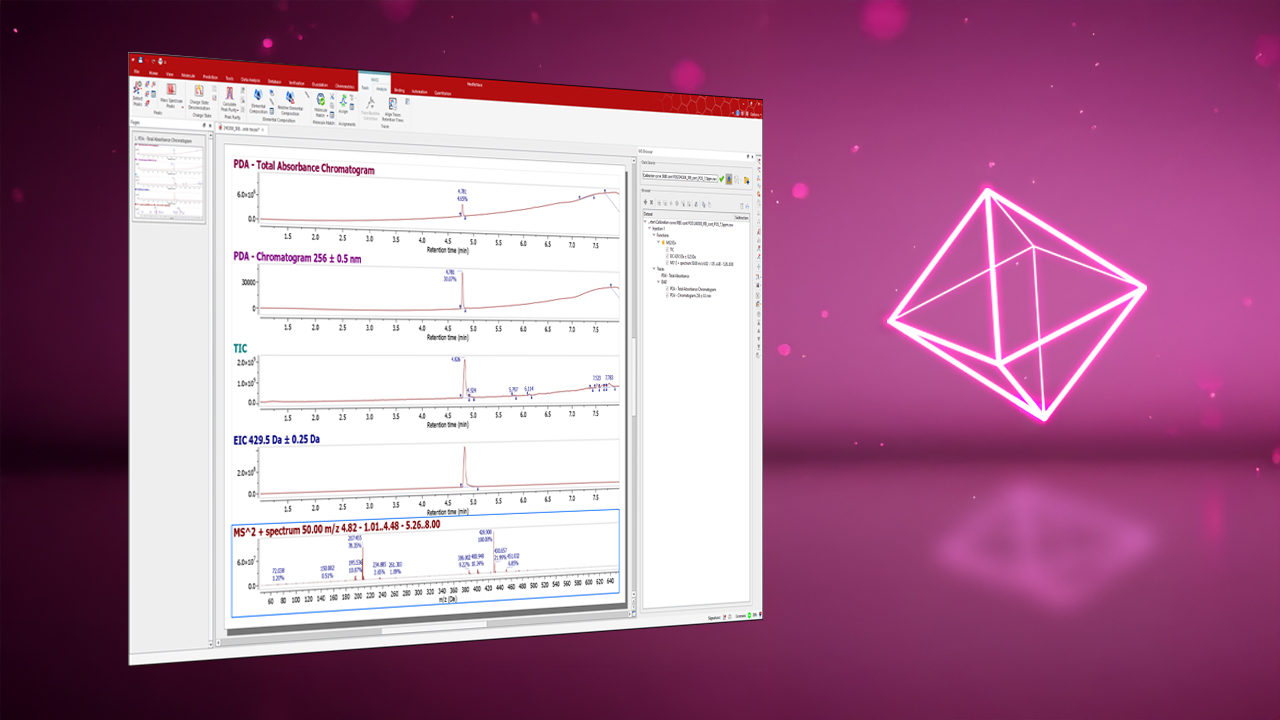
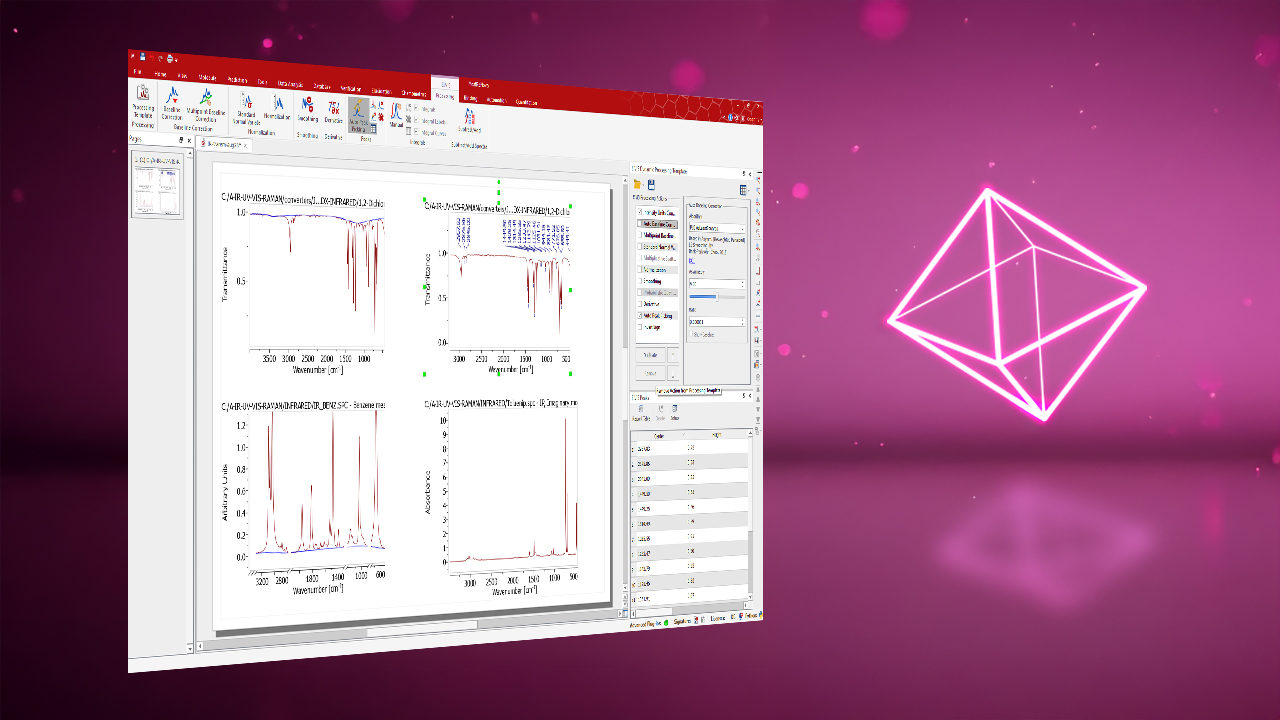
Integrates all analytical data into a single software environment for streamlined handling, analysis, and reporting. View, process, analyze, and report NMR, LC/GC-MS, IR, UV-Vis, IR (FTIR), and Raman data in one intuitive platform. Fewer tools, fewer errors, less training overhead.
Vendor and OS Independence
Compatible with most common NMR and LC/GC-MS formats and fully supported on Windows, macOS, and Linux. Works across instruments and manufacturers without lock-in, ensuring long-term flexibility and system independence.
Multiple Applications
Supports a wide range of workflows including structure verification, mixture analysis, quantification, purity determination, prediction, and advanced analytical applications—adapting to both routine and complex laboratory needs.
Modular and Scalable Architecture
Implement what you need today and expand as your lab grows. Mnova scales seamlessly from single users to global organizations, supporting evolving workflows without system complexity.
Automation and Scripting
No-code automation tools, templates, and scripting support enable standardized workflows and real productivity gains. Fully customizable automation delivers faster processes, fewer errors, and improved reproducibility.
Minimal Learning Curve
An intuitive, user-friendly interface that is easy to learn and widely adopted in academia and industry. Reduces training time and eliminates the need for multiple software applications.
Pre-Packaged Mnova Combos
Mnova’s pre-packaged combos bundle the most-used modules into cost-efficient solutions that deliver more capability for less money. You get the tools labs actually need making it the smartest way to maximize value per license.
Publications
- Ndukwe, I.E., Elipe, M.V.S., Quasdorf, K., Navarro-Vázquez, A., Mardirossian, N., Khachatrian, D., Roosen, P.C. and Smith, A., Anisotropic Nuclear Magnetic Resonance Spectroscopy and Density Functional Theory Methodologies Combine with CASE-3D Analysis for Unambiguous Diastereomeric Differentiation of AMG 176 Macrocycles. ChemistryEurope, 202500074 (2025). https://doi.org/10.1002/ceur.202500074
- Cui-Mei Liu, Wei Jia, Zhen-Dong Hua Automated rapid screening of new psychoactive substances using benchtop NMR technology combined with database search, Forensic Science International 375, (2025). https://doi.org/10.1016/j.forsciint.2025.112564
- D. Williamson, S. Ponte, I. Iglesias, N. Tonge, C. Cobas, E.K. Kemsley Chemical shift prediction in 13C NMR spectroscopy using ensembles of message passing neural networks (MPNNs) Journal of Magnetic Resonance, Volume 368, (2024). https://doi.org/10.1016/j.jmr.2024.107795
- Cobas, C., García-Pulido, J. A., Mora, P., Selva, G., Sykora, S. A New qNMR Compliant Savitzky-Golay Apodization Function for Resolution Enhancement. Magnetic Resonance in Chemistry 63, 90-97 (2025). https://doi.org/10.1002/mrc.5492
- Ossorio-Castillo, J., Rodríguez-Coello, A., Simulating NMR Spectra with a Quantum Computer. arXiv preprint arXiv:2410.20836, (2024). https://arxiv.org/abs/2410.20836
- Costa, P.M., Lysak, D.H., Soong, R., Ronda, K., Wolff, W. W., Downey, K., Steiner, K., Moxley-Paquette, V., Pellizzari, J., Anklin, C., Sharman, G., Cobas, C., Domínguez, S., Jobst, K. J., Cahill, L., Simpson, M. J., & Simpson, A. J. Development of a Simple Cost Effective Oxygenation System for In Vivo Solution State NMR in 10 mm NMR Tubes. Analytical Chemistry 96, 12667-12675 (2024). https://doi.org/10.1021/acs.analchem.4c01390
- Pérez Varela, I., Shear, G., Cobas, C. Molecular Melodies: Unraveling the Hidden Harmonies of NMR Spectroscopy. Molecules 29, 762 (2024). https://doi.org/10.3390/molecules29040762
- Góñez, K. V., García, J. S., Sardina, F. J., Pazos, Y., Saá, Á., Martín−Pastor, M. J-filter: An experiment to simplify and isolate specific signals in 1H NMR spectra of complex mixtures based on scalar coupling constants. Magnetic Resonance in Chemistry 61, 615 (2023). https://doi.org/10.1002/mrc.5396
- Kuhn, S., Cobas, C., Barba, A., Colreavy-Donnelly, S., Caraffini, F., Borges, R. M. Direct deduction of chemical class from NMR spectra. Journal of Magnetic Resonance 348, 107381(2023). https://doi.org/10.1016/j.jmr.2023.107381
- Peng, C., Namanja, A. T., Munoz, E., Wu, H., Frederick, T. E., Maestre-Martinez, M., Iglesias Fernandez, I., Sun, Q., Cobas, C., Sun, C., Petros, A. M. Efficiently driving protein-based fragment screening and lead discovery using two-dimensional NMR. Journal of Biomolecular NMR 77, 39-53 (2023). https://doi.org/10.1007/s10858-022-00410-3
- Soong, R., Downey, K., Moser, A., Monje, P., Jenne, A., Ghosh Biswas, R., Bastawrous, M., Majumdar, R., Lysak, D. H., Adamo, A., Goerling, B., Decker, V., Busse, F., Dominguez, S., Sauer, E., Mikhaylichenko, S., Luk, V., Simpson, A. J. A CASE (Computer-Assisted Structure Elucidation) for Bench-Top NMR Systems in the Undergraduate Laboratory for De Novo Structure Determination: How Well Can We Do?. Journal of Chemical Education 99, 3780-3788 (2022). https://doi.org/10.1021/acs.jchemed.2c00475
- Lumley, J. A., Sharman, G., Wilkin, T., Hirst, M., Cobas, C., Goebel, M. A KNIME Workflow for Automated Structure Verification. SLAS DISCOVERY: Advancing the Science of Drug Discovery 25, 950-956 (2020). https://doi.org/10.1177/2472555220907091
- Cobas, C. NMR signal processing, prediction, and structure verification with machine learning techniques. Magnetic Resonance in Chemistry 58, 512-519 (2020). https://doi.org/10.1002/mrc.4989
- Gallo, V., Ragone, R., Musio, B. et al. A Contribution to the Harmonization of Non-targeted NMR Methods for Data-Driven Food Authenticity Assessment. Food Analytical Methods 13, 530–541 (2020). https://doi.org/10.1007/s12161-019-01664-8
- Barba, A., Dominguez, S., Cobas, C., Martinsen, D. P., Romain, C., Rzepa, H. S., Seoane, F. Workflows Allowing Creation of Journal Article Supporting Information and Findable, Accessible, Interoperable, and Reusable (FAIR)-Enabled Publication of Spectroscopic Data. ACS Omega 4, 3280-3286 (2019). https://doi.org/10.1021/acsomega.8b03005
Curious about our products or services? Let us provide you with the answers you need. Contact us today and discover how we can cater to your needs effectively.