Mnova NMR
About Mnova NMR
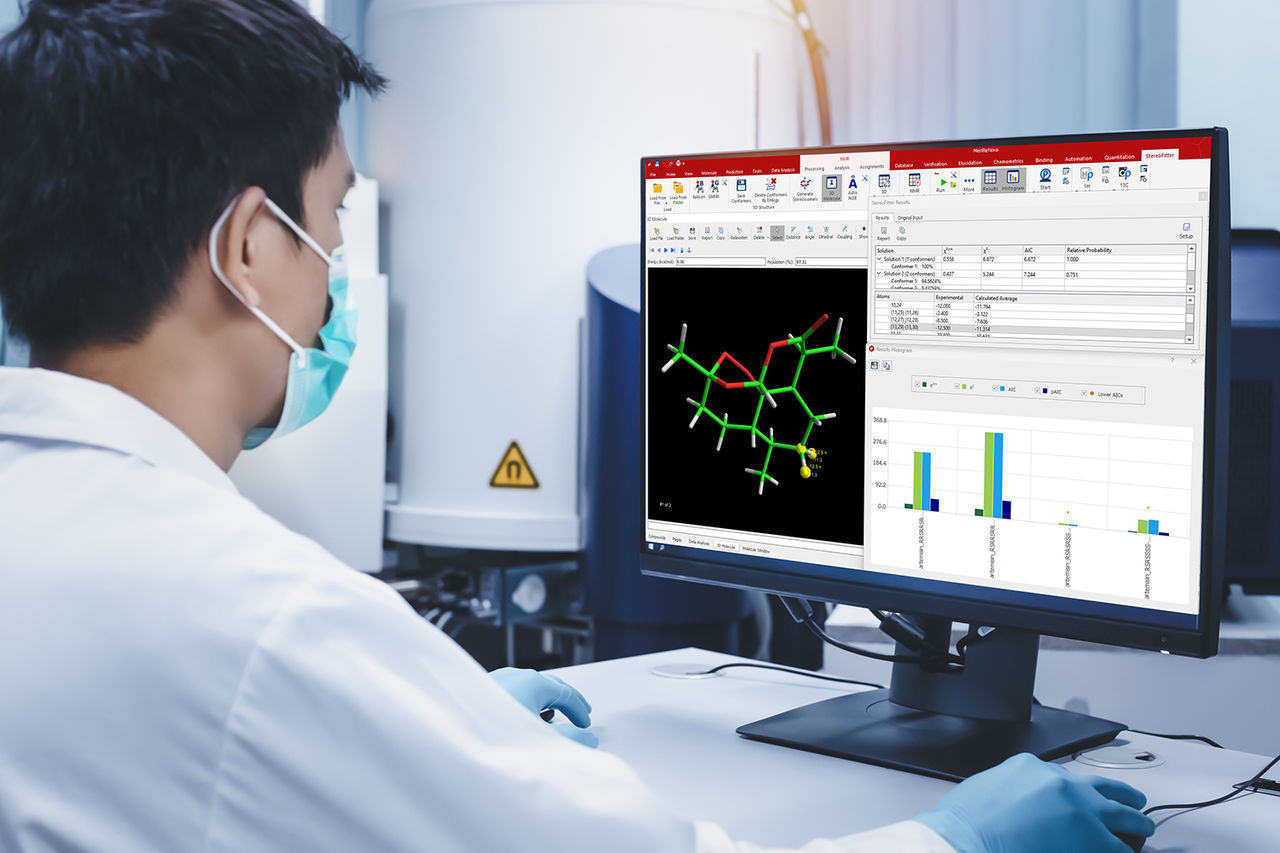
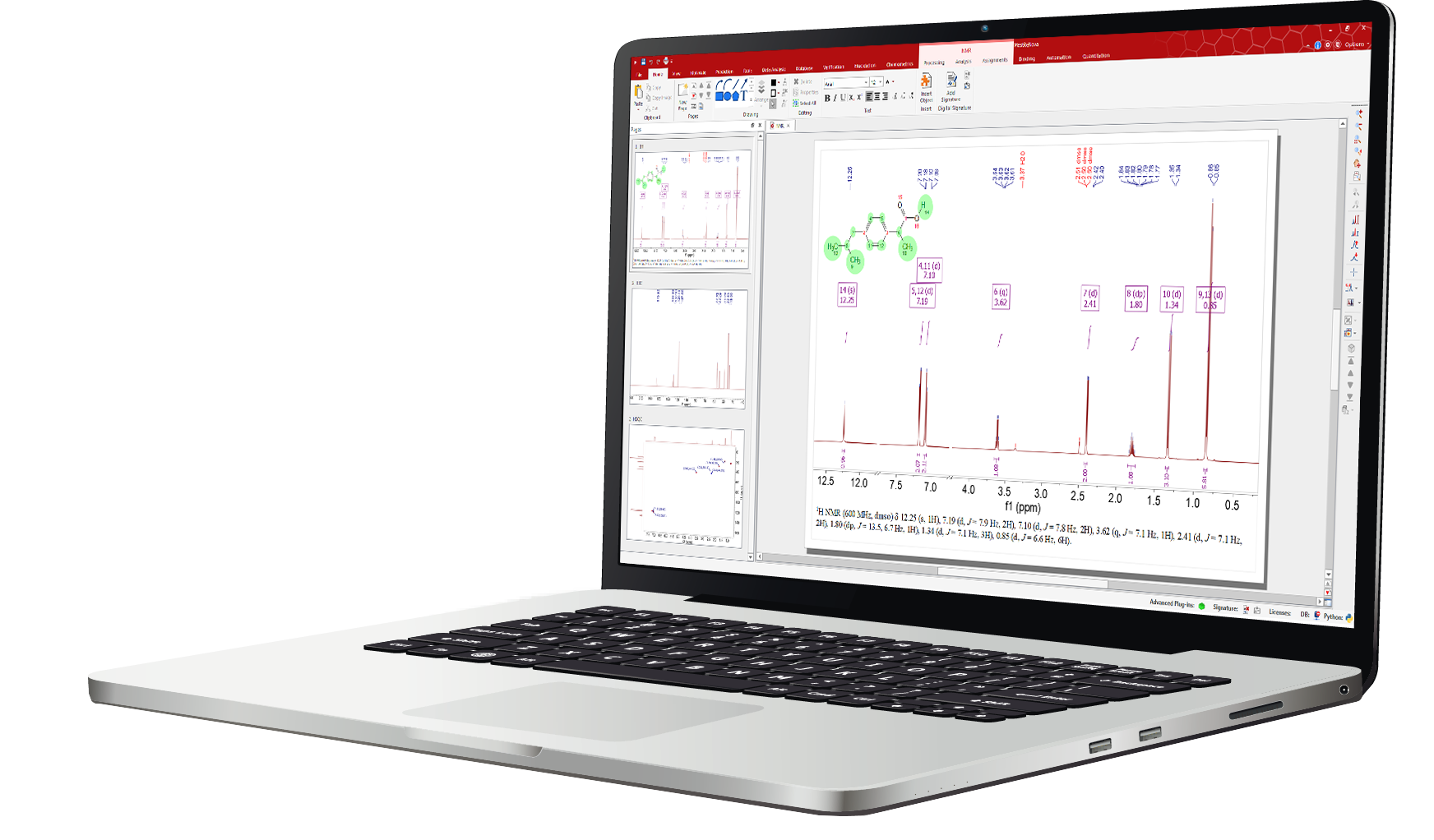
Mnova NMR is a software tool designed for processing, analyzing, and interpreting NMR spectroscopy data. It provides an intuitive interface with advanced tools for spectral analysis and reporting across a wide range of NMR experiments and file formats. Mnova NMR integrates seamlessly with multiple plugins for applications such as reaction monitoring, qNMR, mixture analysis, structure elucidation, verification, and fragment-based drug discovery.
Data Insights
Features
Comprehensive 1D and 2D NMR Data Analysis
3D NMR Data Handling and Visualization Capabilities
Best-in-Class NMR Spectral Processing and Analysis Tools
State-of-the-Art Algorithms for AI-Powered Peak Picking
Assignments Tool
Multi-Vendor NMR Integration
Mnova is a versatile, cross-platform software package compatible with Windows, macOS, and Linux, operating without hardware restrictions. Mnova NMR is fully vendor-independent, designed for heterogeneous environments, and supports datasets from all major NMR spectrometer manufacturers, including Bruker, Agilent/Varian, Jeol, Magritek, Oxford Instruments, and Nanalysis.
Check out the full list of supported vendors and NMR file formats. For any questions regarding your datasets format compatibility contact us!
Read MoreEasy Handling of Multiple Spectra
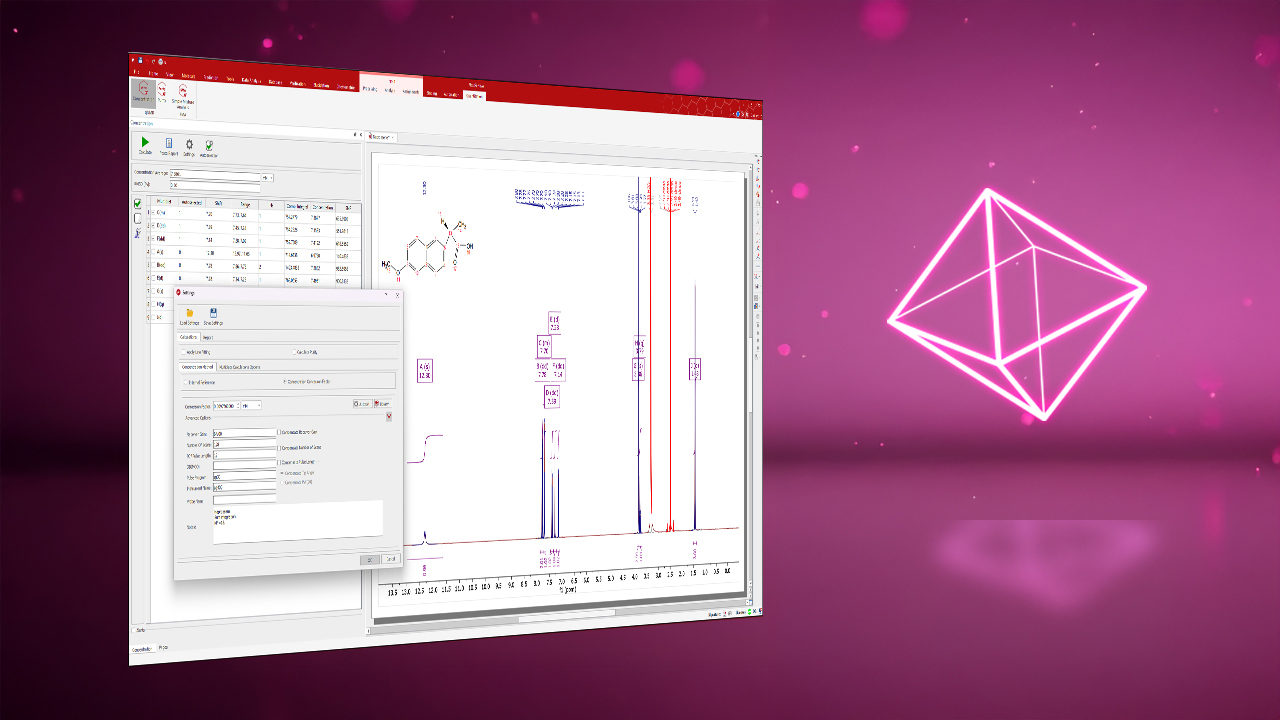
The Mnova NMR graphical interface enables fast, intuitive interaction with multiple 1D and 2D NMR spectra, offering flexible visualization, processing, and analysis tools. It allows:
- Stacking and superimposing multiple 1D or 2D spectra
- Simultaneous processing, peak picking, and integration across multiple spectra
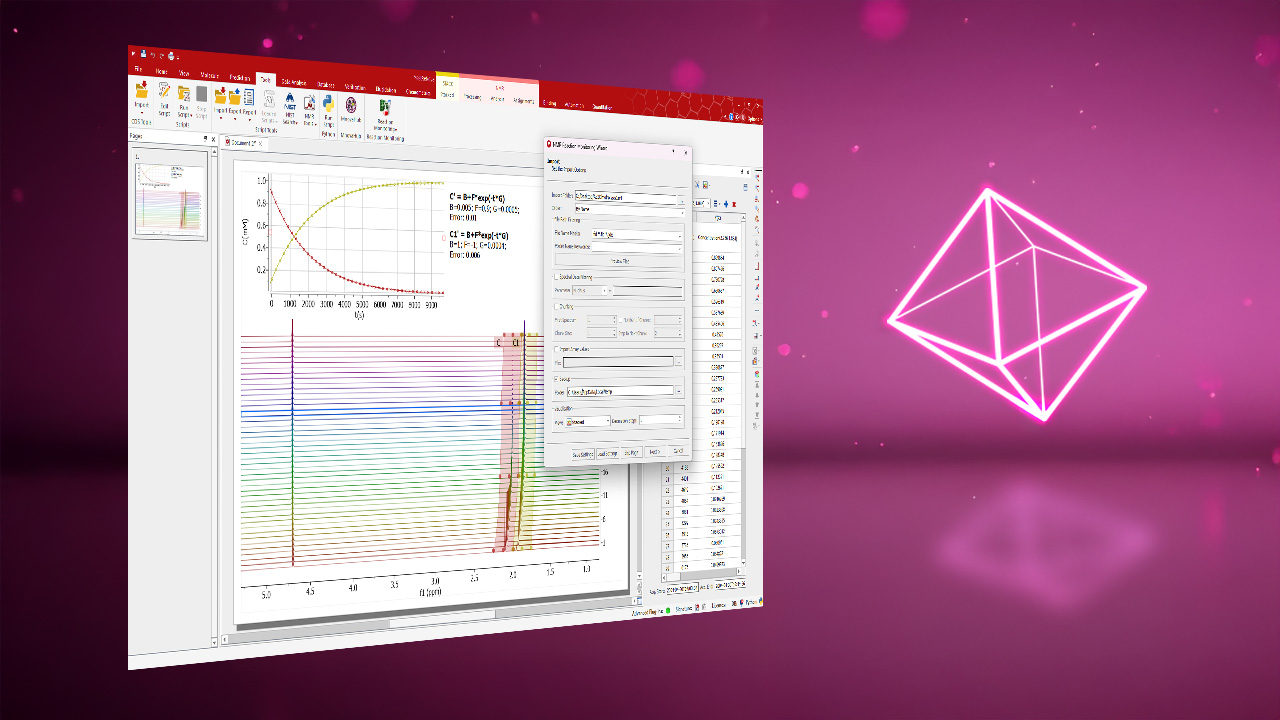
- Curve fitting to peak parameters for related spectra (e.g., reaction monitoring, relaxation studies, DOSY processing).
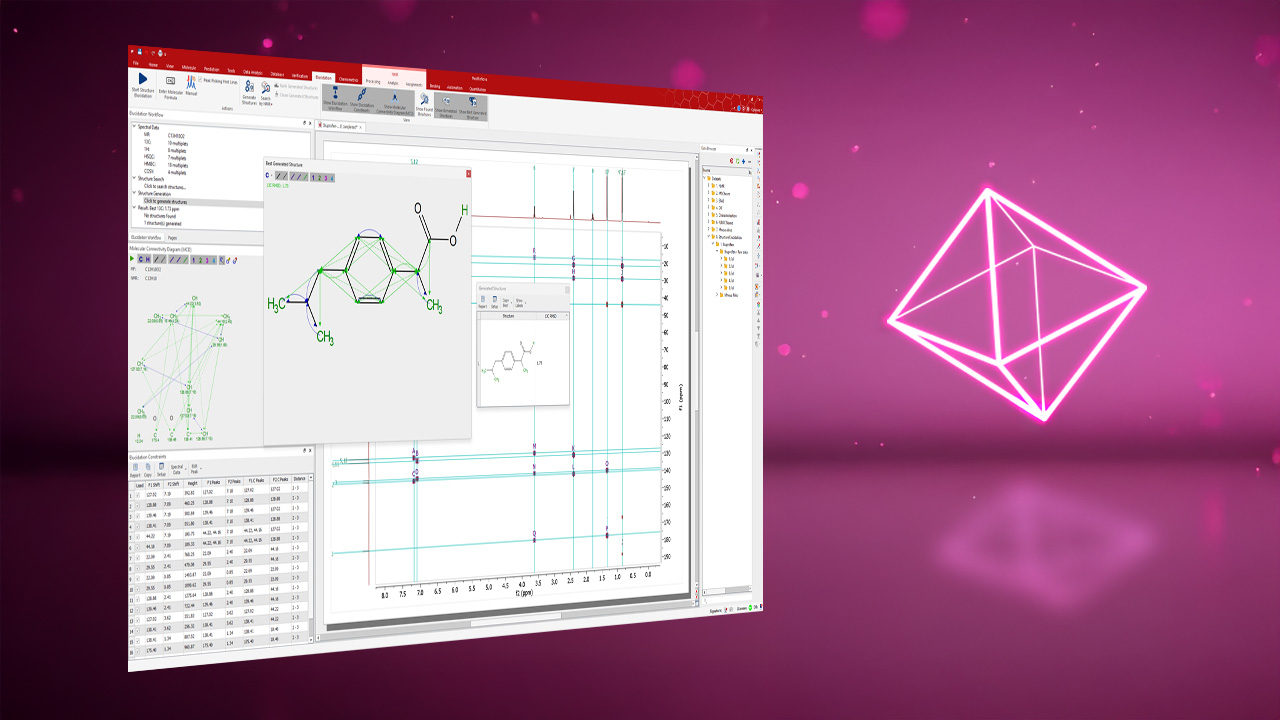
Data Analysis to Calculate NMR-Related Molecular Information
13C/HSQC Molecular Search
DOSY Module
Easy Access to NMR Data from Different Repositories
Benefits
All-in-One NMR Analysis: Analyze all your NMR data in a single, consistent environment. No need for multiple software tools, making workflows simpler, faster, and more reliable.
Intuitive and Continuously Evolving: Mnova is designed to support users from academia to industry, adapting to diverse environments, workflows and evolving research needs.
Reduced Learning Curve: Intuitive design allows users to focus on extracting insights from NMR data rather than struggling to learn the software.
Advanced Applications: Built on top of powerful standard processing features, Mnova offers tools to support a broad range of NMR applications, such as reaction monitoring, qNMR, mixture analysis, chemometrics, fragment-based drug discovery, structure elucidation, verification, and biologics.
Time-Saving for Scientific Focus: Automating data processing, analysis, and reporting frees researchers to concentrate on high-value scientific activities, rather than spending time on routine tasks.
More Information
Publications
1. Cobas, C., García-Pulido, J. A., Mora, P., Selva, G., Sykora, S. A New qNMR Compliant Savitzky-Golay Apodization Function for Resolution Enhancement. Magnetic Resonance in Chemistry 63, 90-97 (2025). https://doi.org/10.1002/mrc.5492
2. Williamson, D., Ponte, S., Iglesias, I., Cobas, C., Tonge, N. M., & Kemsley, E. K. (2024). Chemical shift prediction in ¹³C NMR spectroscopy using ensembles of message passing neural networks (MPNNs). Journal of Magnetic Resonance, 368, 107795 https://doi.org/10.1016/j.jmr.2024.107795
3. Costa, P.M., Lysak, D.H., Soong, R., Ronda, K., Wolff, W. W., Downey, K., Steiner, K., Moxley-Paquette, V., Pellizzari, J., Anklin, C., Sharman, G., Cobas, C., Domínguez, S., Jobst, K. J., Cahill, L., Simpson, M. J., & Simpson, A. J. Development of a Simple Cost Effective Oxygenation System for In Vivo Solution State NMR in 10 mm NMR Tubes. Analytical Chemistry 96, 12667-12675 (2024). https://doi.org/10.1021/acs.analchem.4c01390
4. Pérez Varela, I., Shear, G., Cobas, C. Molecular Melodies: Unraveling the Hidden Harmonies of NMR Spectroscopy. Molecules 29, 762 (2024). https://doi.org/10.3390/molecules29040762
5. Góñez, K. V., García, J. S., Sardina, F. J., Pazos, Y., Saá, Á., Martín−Pastor, M. J-filter: An experiment to simplify and isolate specific signals in 1H NMR spectra of complex mixtures based on scalar coupling constants. Magnetic Resonance in Chemistry 61, 615 (2023). https://doi.org/10.1002/mrc.5396
6. Kuhn, S., Cobas, C., Barba, A., Colreavy-Donnelly, S., Caraffini, F., Borges, R. M. Direct deduction of chemical class from NMR spectra. Journal of Magnetic Resonance 348, 107381(2023). https://doi.org/10.1016/j.jmr.2023.107381
7. Soong, R., Downey, K., Moser, A., Monje, P., Jenne, A., Ghosh Biswas, R., Bastawrous, M., Majumdar, R., Lysak, D. H., Adamo, A., Goerling, B., Decker, V., Busse, F., Dominguez, S., Sauer, E., Mikhaylichenko, S., Luk, V., Simpson, A. J. A CASE (Computer-Assisted Structure Elucidation) for Bench-Top NMR Systems in the Undergraduate Laboratory for De Novo Structure Determination: How Well Can We Do?. Journal of Chemical Education 99, 3780-3788 (2022). https://doi.org/10.1021/acs.jchemed.2c00475
8. Cobas, C. NMR signal processing, prediction, and structure verification with machine learning techniques. Magnetic Resonance in Chemistry 58, 512-519 (2020). https://doi.org/10.1002/mrc.4989
9. Gallo, V., Ragone, R., Musio, B. et al. A Contribution to the Harmonization of Non-targeted NMR Methods for Data-Driven Food Authenticity Assessment. Food Analytical Methods 13, 530–541 (2020). https://doi.org/10.1007/s12161-019-01664-8
Curious about our products or services? Let us provide you with the answers you need. Contact us today and discover how we can cater to your needs effectively.